|
Information content for DNA is by codons, i.e., per triplets of bases. 
Comparison of information content in DNA and protein sequences at PAM distances from zero to 100 PAMs. Therefore, the codon redundancy can explain the information loss.įigure 1. For example, all the four codons GCT, GCC, GCA, GCG code for a single amino acid Alanine. Where did the information get lost? Let's see, one codon of three bases could potentially make 64 different amino acids (4^3=64), but only 20 of them are in use, resulting in most of the codons being redundant, in other words, coding for same amino acids. However, each amino acid can only carry about 4.3 bits.Ĭonsequently, there is a difference of about 1.7 bits less information in protein sequences. So, it looks like protein sequences could have more information than DNA sequences, but wait, DNA codes for protein using codons consisting of three bases each Thus, DNA uses six bits of information (3x2=6) to code one amino acid. Protein sequences consist of 20 different amino acids, and we can calculate the maximum amount of information each amino acid can carry the same way as for DNA by taking the logarithm of base two of 20 which gives ~4.3 bits. DNA sequence consists of four different bases, A, T, G, C and we can calculate the maximum amount of information each base can carry by taking the logarithm of base two of four which gives two bits. To be able to construct custom scoring schemes for DNA sequence alignments.ĭNA Sequence Alignment vs Protein Sequence Alignmentīefore we dig into DNA scoring matrices, let's compare the amount of information that protein and DNA sequences can carry, because, interestingly protein and DNA sequences contain differing amounts of information. 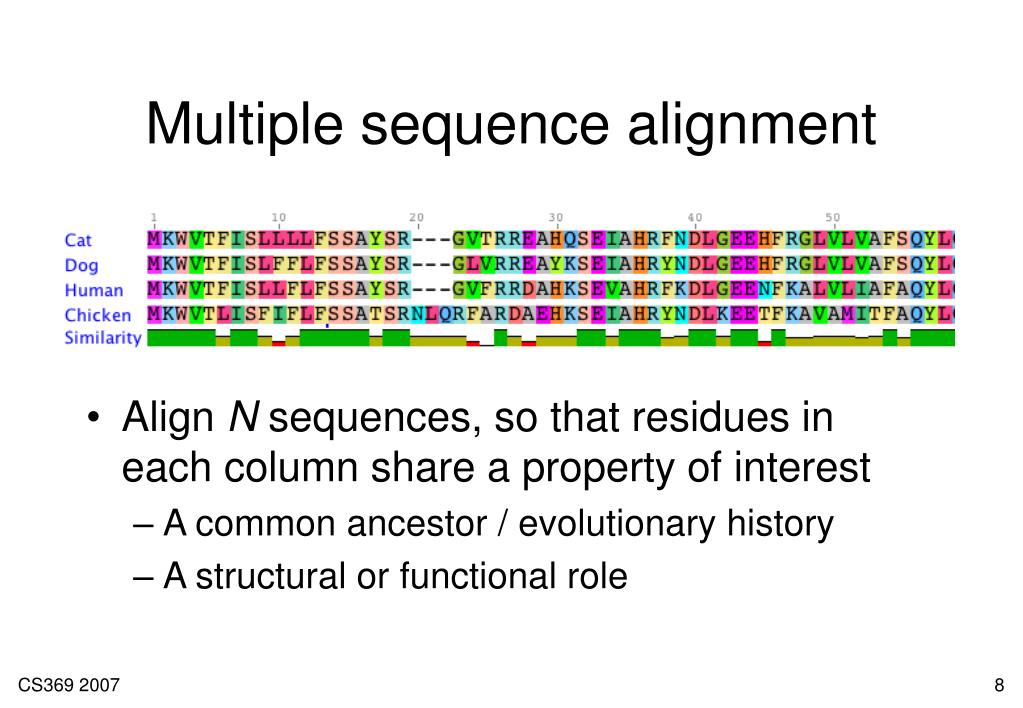
To provide a basic understanding of various scoring schemes' effect on sequence alignments.Ģ.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
